Quick instruction for setting up standalone BLAST search
MAC OS 10.6.8, Terminal "bash"
1. Go to NCBI website (Note: Safari has problem with
downloading database files, use FireFox instead)
2. Click "NCBI FTP Site". You can find the link at bottom right on the
page. (Note: select "Guest" in case you are asked for user ID and
password)
3. Go to “BLAST Basic Local Alignment Search Tool” and then
click the link. You will see list of
files and folders.
4. Open a folder "executables", go to
"LATEST", and then double click "ncbi-blast-2.2.26+-universal-macosx.tar.gz"
or latest version. Download starts
automatically. You can find the blast
folder in the folder “Downloads”
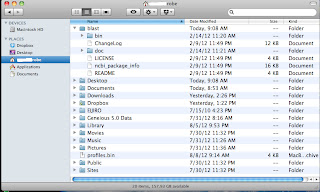
5. Rename the downloaded folder to "blast", and
then move it into "Home Directly" (the one with the icon of house,
where folders “Desktop”, “Documents”, “Downloads” are stored).
6. Before setting up stand alone blast, let’s review some
basic commands in Linux
Start "Terminal" and then try following command:
Note: You can find the program “Terminal” in “Utilities”
folder in “Applications”
Show your current directly (folder)
pwd
List all the files in a directly (folder) (hidden files
included)
ls -a
Move to a different folder (change directly)
cd [folder name]
cd [folder]/[folder]
Back to Home Directly
cd
Create or edit a file
vi [file_name]
example) vi .bash_profile
type "a" to enter
"edit mode"
hit "esc" to exit edit
mode
hit "ZZ" (shift+z twice)
to save and back to normal window
Delete an existing file (remove)
rm [file_name]
example) rm .bash_profile
Look content of a file
more [file_name]
example) more output.txt
(hit "q" to quit the
command)
7.
Create .bash_profile (Look Step 6) in your “Home Directory”. Here are commands to create a new file for setting a PATH.
vi .bash_profile [hit enter]
(this creates a file named “.bash_profile”)
[hit "a"] (this
is to enter edit mode)
export PATH=/Users/[name of your home directly]/blast/bin:$PATH
[hit "esc"] (exit edit mode)
[hit "ZZ"]
(save file and then back to terminal)
Restart Terminal (this is needed to activate the path).
Note: Why we need to set PATH?
The program "Terminal" looks for command programs
stored in certain folders (folders can be visualized by typing
"$PATH"). All the commands
listed above (ls, more) are stored as program files in one of the folders
listed below.
/Users/tomokurobe/blast/bin:/usr/bin:/bin:/usr/sbin:/sbin:/usr/local/bin:/usr/X11/bin:
No such file or directory
This means...
Terminal looks following folders to find programs:
/Users/tomokurobe/blast/bin
/usr/bin
/bin
/usr/sbin
/sbin
/usr/local/bin
/usr/X11/bin
or return a message "No
such file or directory"
In the default setting, "Terminal" doesn't look
the newly added folder "blast", so that we need to set PATH. Otherwise Terminal cannot find blast programs
stored in the new folder. In worst case,
we can move blast programs in one of the folders above, but not recommended as
you will be asked for password every time you make change. Those folders contain important files for
running OSX so it’s designed that users cannot make changes easily.
8.
8-1) Go to the NCBI FTP site and download database file in
FASTA format (double click fasta file you want to use). For test purpose, choose small database like
"yeast.aa". "nr" database is large, even the compressed file is
bigger than 10 Gb. It takes very long
time to run search program. For this
tutorial, you can download fasta file for “yeast.aa”
8-2) Now fasta file is downloaded in folder
“Downloads”. Create a new folder
"db" in "blast" folder and then move the FASTA file
“yeast.aa” in it.
8-3) In Terminal, type “pwd” to see your current location. You should be in your Home Directly. Type “ls” and make sure that there is a
folder “blast”. Type "cd
blast/db" to reach the database folder.
Type “ls” to see there is the fasta file, “yeast.aa”.
8-4) To format database, type following command
makeblastdb -in yeast.aa -dbtype prot -parse_seqids (-parse_seqids is not necessary!)
This command creates 8 files (.phr .pin .pnd .pni .pog .psd
.psi .psq). All the files are needed for
running blast search.
9. Run blast search
Note: we don't have to specify the database file created in
the previous step. Simply type database
name and don't add any file extension.
Here is the instruction.
9-1) Prepare test sequence(s) in fasta format and then name it
“test.fasta”
9-2) Stay in the "db" folder and then type in the
following command:
blastp -query test.fasta -db yeast.aa -out output.txt
(Note: this command creates output file “output.txt” and you see result in regular output format, 50 hits)
more test.txt (to look your result)
blastp -query test.fasta -db yeast.aa -out output.txt
-max_target_seqs 1 -outfmt '7 std sseqid
sgi'
(Note: tubular format with comments, 1 hit. -outfmt ‘[number] [std(standard)] [other info
you want to add]’)
blastp -query test.fasta -db yeast.aa -out output.txt
-max_target_seqs 1 -outfmt '6 sgi'
(Note: tubular format without comments, 1 hit)
Important Note: Comment from NCBI staff "Be careful using -max_target_seqs 1. You could miss the best hit. I would test with a higher number, like 50, to be sure setting to 1 gives the top hit with your query and db"
10. Retrieve definition of gene (gene name). This can be done just adding stitle in outfmt option.




No comments:
Post a Comment